This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
|
What is Gene Ontology? [1,2]
The Gene Ontology (GO) project is an effort in the scientific community to streamline descriptions of genes and their products into a universal system in different databases. The benefit of using one common language between all academic groups is that it will be more effective and less time consuming to search certain genes. In the past a wide range of terminology was used to describe genes which often times would lead to confusion when attempting to search key terms or to identify functionally equivalent terminology. What originally started off only being used by a few databases now encompasses many including FlyBase, Mouse Genome Database, and Saccharomyces Genome Database. Three different domains have been made that help describe genes and their products. This controlled vocabulary can be viewed from different levels: from a more general observation/description or down to more specific level. The three different domains of gene ontology are cellular components, molecular functions, and biological processes. |
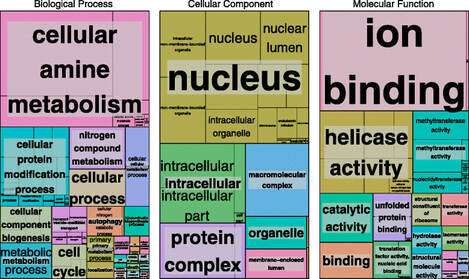
The Three Domains of Gene Ontology
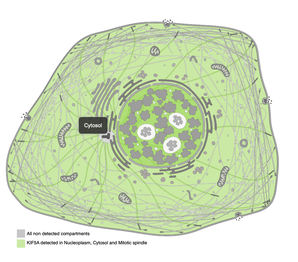
Cellular ComponentDefines the location in which the gene product functions in a cell. This can be compartments or different organelles within the cell. This domain only focuses on the anatomy of the cell.
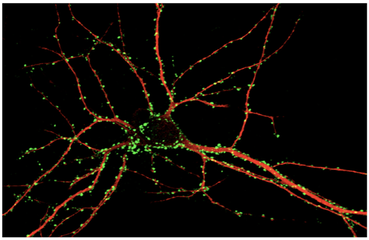
In terms of KIF5A, the protein of this gene is localized in the cytosol and the nucleoplasm as well as at the mitotic spindle of cells. This protein is typically found in neurons and near complexes that aid in transportation. Other cellular GO terms used to describe the gene are: neuronal cell body, culinary rootlet, dendrite cytoplasm, kinesin complex, microtubule, and membrane. |
Biological ProcessesDefines the larger processes accomplished by the multiple cellular activities. This can be biological processes that maintain the cell or signal transduction pathways.
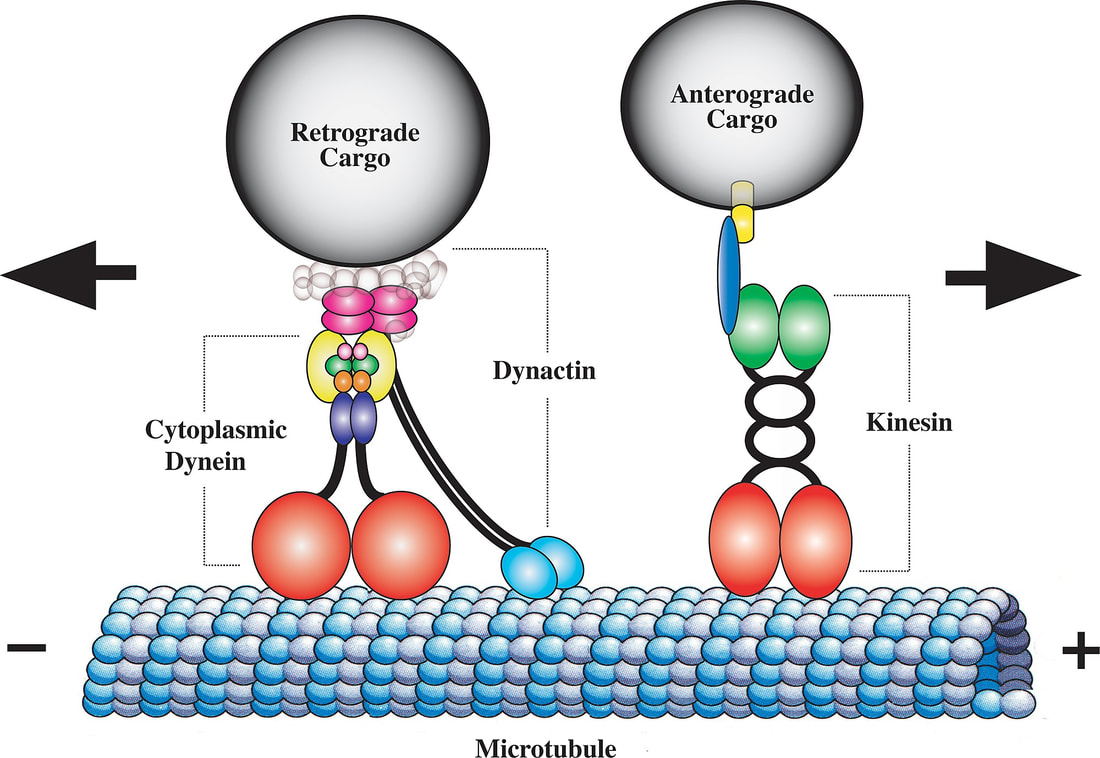
KIF5A plays apart in axonal organelle transport carrying vesicles through the axon retrograde and anterograde. Some GO terms used to define the gene biological function are: anterograde dendritic transport of neurotransmitter complex, chemical synaptic transmission, microtubule based movement, axon guidance, synaptic vesicle transport, vesicle-mediated transport, retrograde vesicle mediated transport (Golgi to endoplasmic reticulum), protein localization |
Molecular FunctionDefines the molecular-level activity that the product of the gene does. These do not describe processes or pathways, but rather the activity that makes them up. Can correspond to activities done by that sole product of the gene or by products of a few genes.
KIF5A works in unison with other proteins to make up a kinesin motor protein that functions by the aid of ATP, pushing cargo along microtubules. Some GO terms used are: ATP binding, protein binding, kinesin binding, microtubule binding, ATPase activity, ATP-dependent microtubule motor activity (plus end directed), motor activity, microtubule motor activity. |
Conclusion
Gene ontology plays an important role in understanding the function of KIF5A. Seeing the different GO terminology associated with the protein illustrates how a defective gene can result in the symptoms found in patients with hereditary spastic paraplegia. The GO terminology shows us that the KIF5A protein can be found in the cytoplasm of cells (especially neurons) and plays a role in axonal organelle transport by being part of a Kinesin motor complex. This type of analysis is useful when looking at any type of gene or protein. As we did with KIF5A specifically here, we are able to see the big picture and how that protein plays a functional role in it.