This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
|
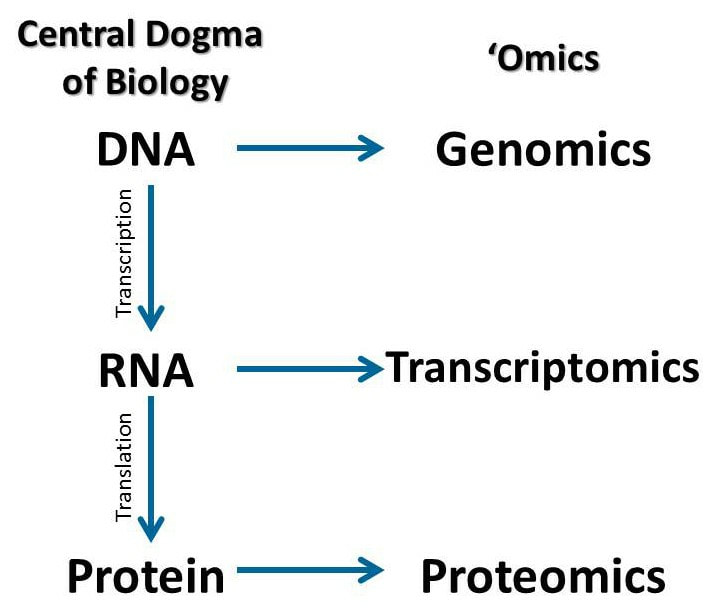
What is the Transcriptome? [1] DNA is typically transcribed with the help of proteins in the cell to produce mRNA. The mRNA transcribed is known as a transcript and the study of the collection of all of these transcripts created based off the DNA is known as the transcriptome. In other words, the transcriptome is the collection of all gene readouts present in the cell produced by the genome. What is Transcriptomics? [1] This area of genetics specifically looks at the expression of genes and analyzes the changes that occur in different cells under various conditions. Every cell contains the same genes, but different cells show different expression levels which is dependent on many factors. Commonly, transcriptomics is looked at to determine the activity of diseased cells versus normal cells. There are two main methods in which transcriptomics is analyzed: microarray analysis and RNA sequencing. |
|
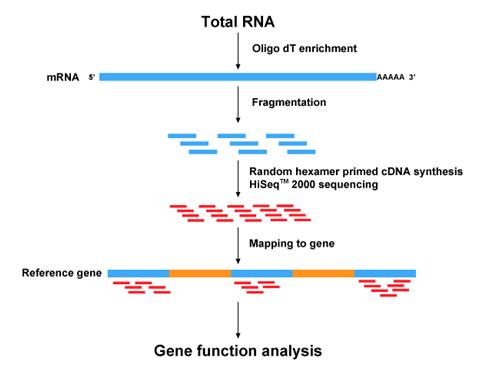
RNA Sequencing: [3] This approach to transcriptome profiling is relatively new and utilizes deep sequencing technologies. In this method, mRNA is converted to cDNA with adaptors at either end of the cDNA. Each sequence is then sequenced in a high throughput manner either using single-end or pair-end sequencing. Any high throughput sequencing system can be utilized to achieve this, however the most common program used is Illumina IG. These sequences are then aligned with a reference transcript or genome . |
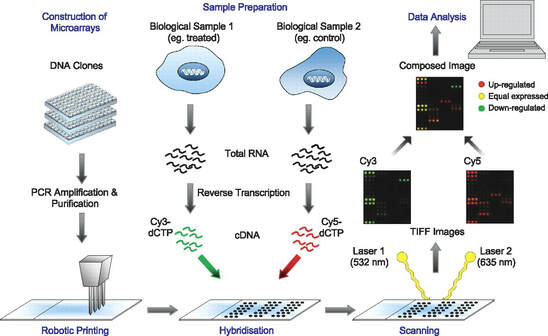
Microarray Analysis: [2] Microarray analysis is a method that uses a microchip with probes and allows the researcher to look at expression levels of multiple genes at once. Each spot that has a probe is a known gene or DNA fragment that has been placed on the chip for observation. DNA samples are labelled and introduced to the slide so that hybridization may occur. The results are then viewed through imaging and computational analysis of the microchip. This will generate a "list" of genes that are expressed in a cell or tissue. |
Conclusion
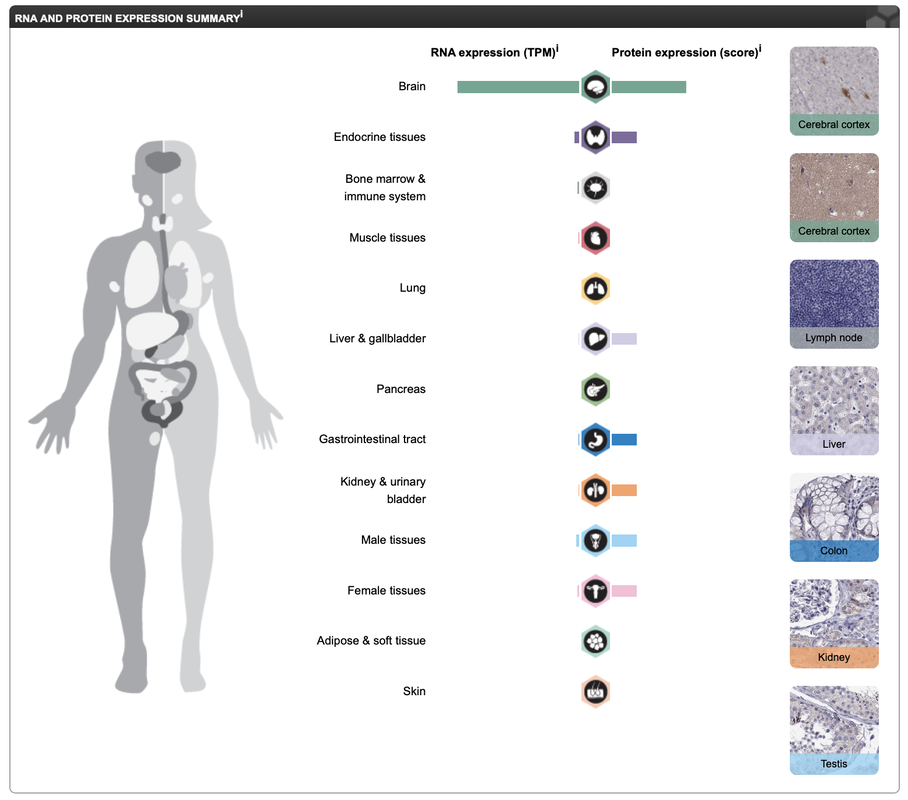
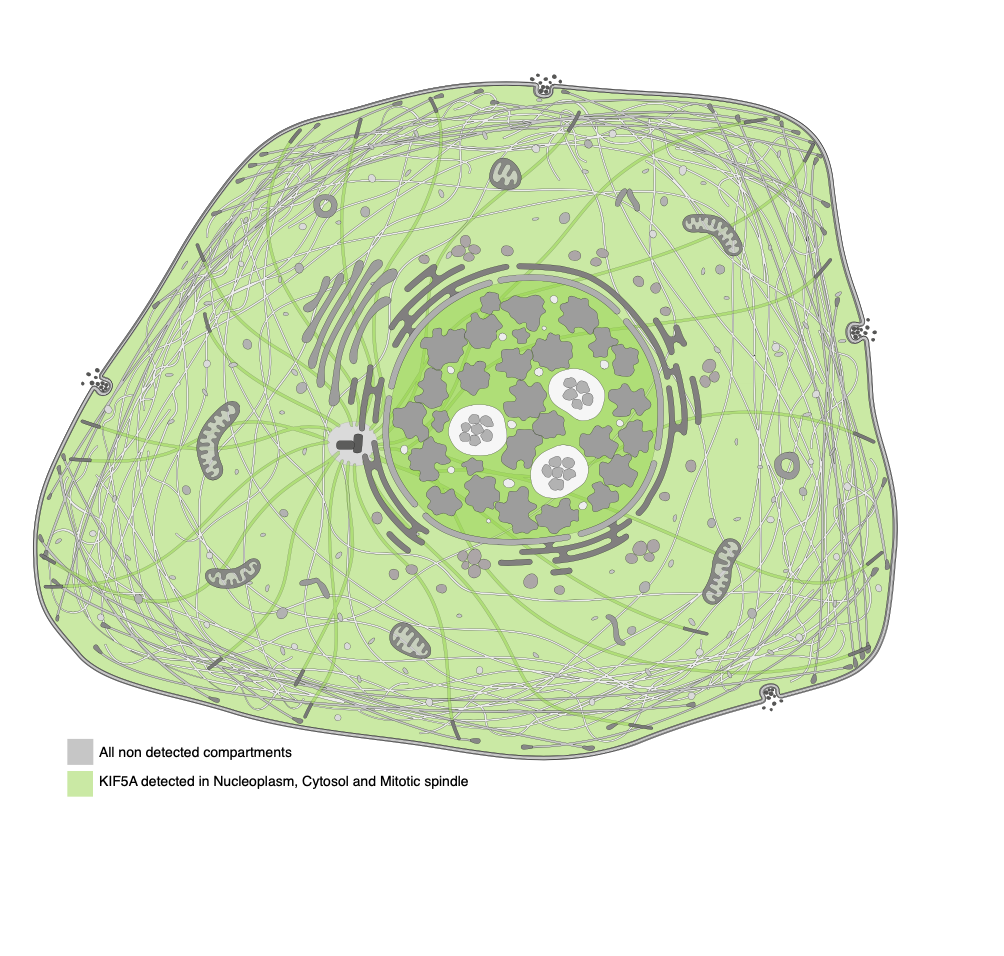
Observing the expression level of KIF5A is useful because it allows us to confirm where the protein localizes with in both cells and tissues. Using the the Human Protein Atlas, a visually appealing database that records gene expression and localization, I was able to observe that KIF5A is mainly found within neurons as well as the brain. In terms of cellular localization, KIF5A is found within the cytoplasm of the cell and especially near mitotic spindle. This information reaffirms the GO classifications (gene ontology page) of KIF5A indicating that the protein plays a crucial role in proper neuronal development.
References:
[1] Transcriptome Fact Sheet (August 27, 2015). Retrieved from https://www.genome.gov/about-genomics/fact-sheets/Transcriptome-Fact-Sheet
[2] DNA Microarray Fact Sheet ( August 27, 2015) Retrieved from https://www.genome.gov/about-genomics/fact-sheets/DNA-Microarray-Technology
[3] Mackenzie, R. ( April 6, 2018). Retrieved from https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461
[1] Transcriptome Fact Sheet (August 27, 2015). Retrieved from https://www.genome.gov/about-genomics/fact-sheets/Transcriptome-Fact-Sheet
[2] DNA Microarray Fact Sheet ( August 27, 2015) Retrieved from https://www.genome.gov/about-genomics/fact-sheets/DNA-Microarray-Technology
[3] Mackenzie, R. ( April 6, 2018). Retrieved from https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461