This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Phylogenetics? [1,2]
|
Phylogenetics is the study of evolutionary relatedness among different groups of organisms and determining the evolutionary history of extant species. This is typically organized in the construct of a phylogeny tree using both morphologies and sequence data. Using this information we can make different conclusions about how different species came to be or and when different speciation events may have occurred.
Three Different Types of Phylogenetic Trees [1,2]
|
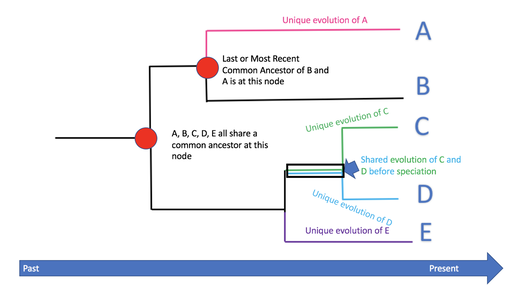
Phylogeny Tree Basics [1,2]
Taxa - set of organisms or groups of organisms Nodes - a point on the phylogeny tree where a single branch breaks into more than one branch Common Ancestor - found where the two branches intersect to form a node Sister Group - two species that come from the same node Outgroup - a taxon outside the group of interest Topology - the branching pattern of the gene Distance Scale - the distance represents the amount of differences between each organism Clade - a group of two or more taxa that includes their common ancestor |
Constructing the Phylogeny Tree for KIF5A
Step 1: Find all protein sequences in FASTA format for homologs and save document as a .txt file. All of the homologs with their prospective FASTA sequences can be found on Homlogene.
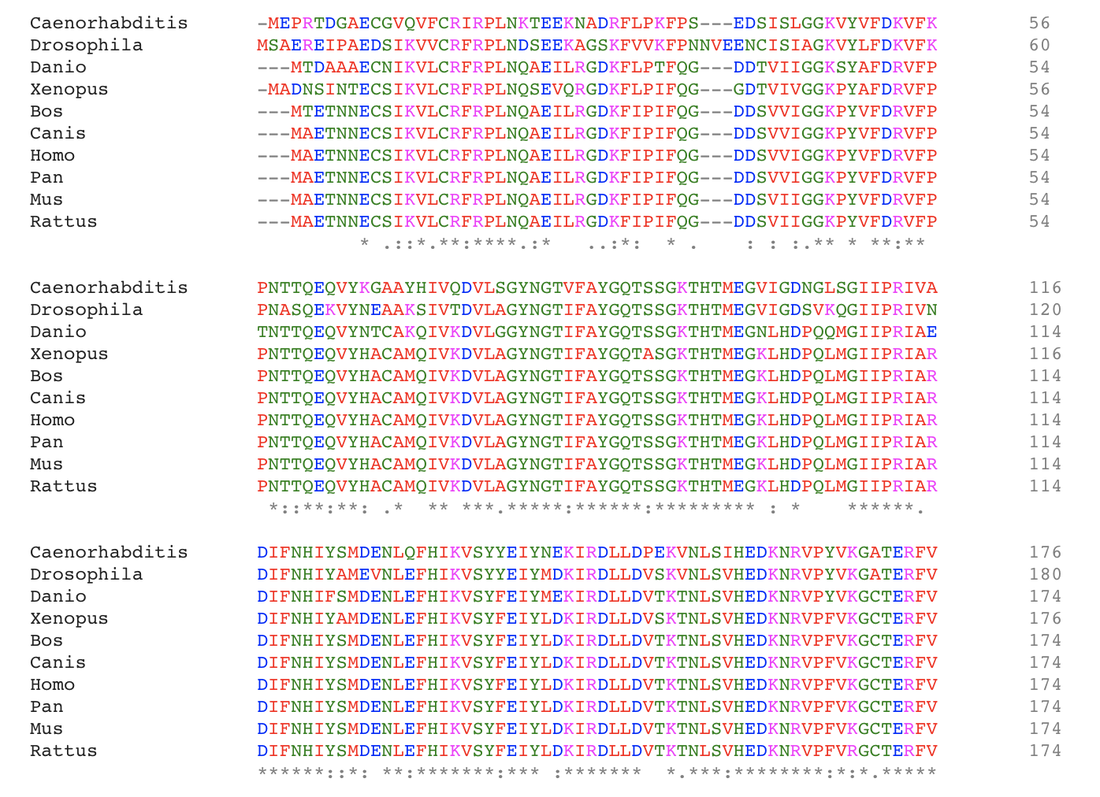
Step 2: With the FASTA data attained previously, align the sequences using Clustal Omega.
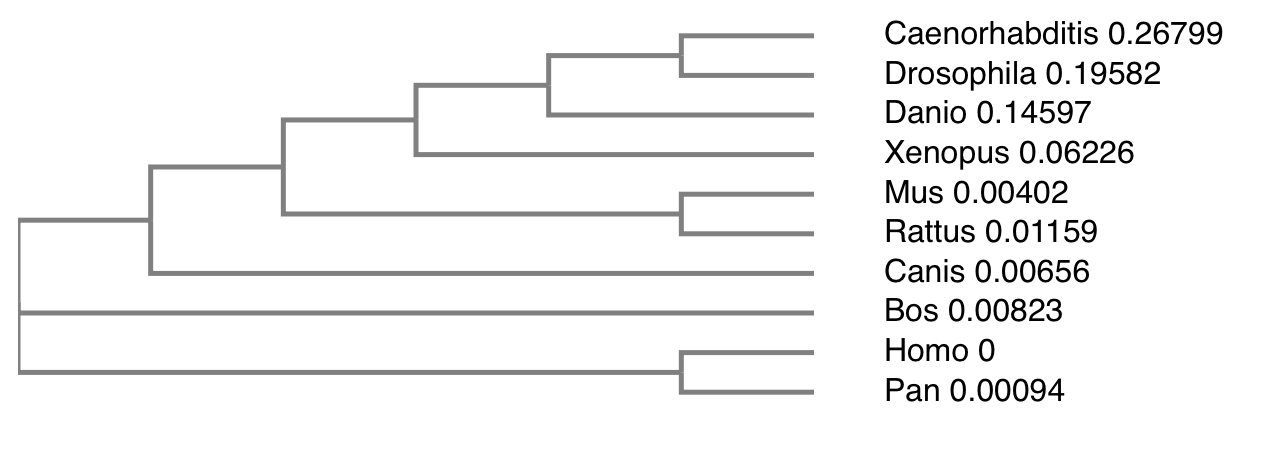
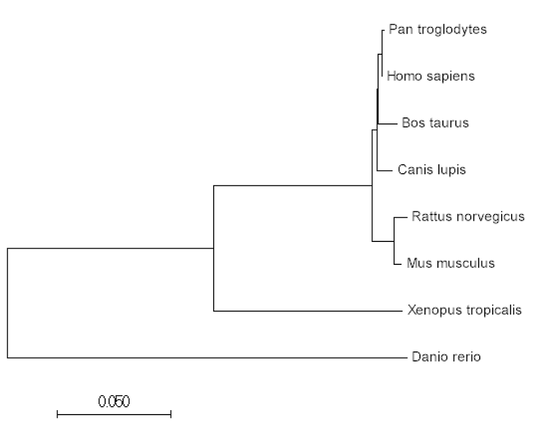
Step 3: Make a simply phylogeny tree on Clustal Omega. This simple phylogeny tree utilizes the Neighbor Joining Method.
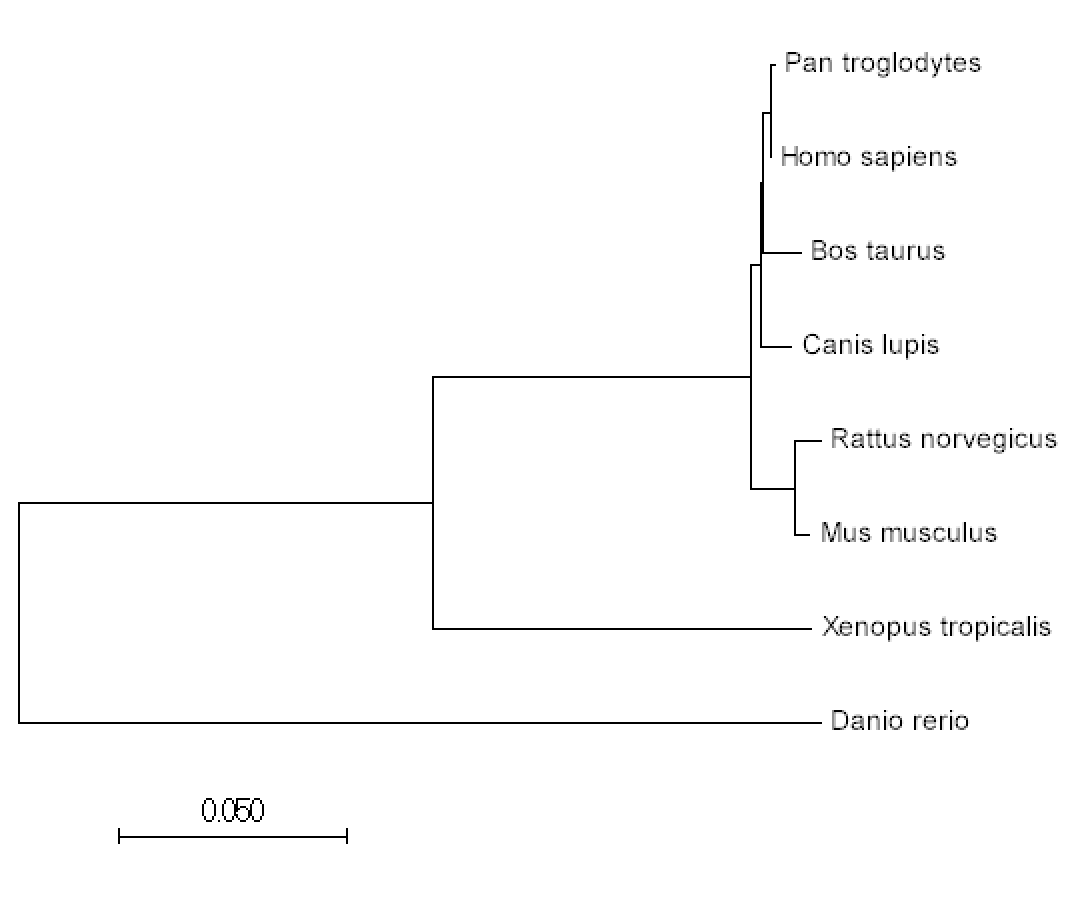
Step 4: Using the FASTA data, create phylogeny trees using the three different methods.
Discussion
Phylogenetic trees help us visualize how conserved KIF5A is amongst other homologs and how the gene may have diverged over time. While identifying homologs through Homologene may have given us an understanding that KIF5a has a conserved function to biological systems of different organisms, phylogenetic trees illuminate the history of the gene and the extent of its relatedness amongst other groups. One thing that is noticeable after the construction of both the maximum likelihood tree and the neighbor joining tree, both have the same organization and layout. As a result, either tree is fair game to choose when considering the evolutionary history of KIF5A.
References:
[1] What is phylogenetics? (2016, June 08). Retrieved from https://www.ebi.ac.uk/training/online/course/introduction-phylogenetics/what-phylogenetics
[2] Chhotwani, M., Francis, S., & Pal, A. (n.d.). Comparison of different phylogenetic tree generation methods. Retrieved from https://cise.ufl.edu/~sarath/BioInformatics/
[1] What is phylogenetics? (2016, June 08). Retrieved from https://www.ebi.ac.uk/training/online/course/introduction-phylogenetics/what-phylogenetics
[2] Chhotwani, M., Francis, S., & Pal, A. (n.d.). Comparison of different phylogenetic tree generation methods. Retrieved from https://cise.ufl.edu/~sarath/BioInformatics/